Structural Determinants of the Dictyostatin Chemotype for Tubulin Binding Affinity and Antitumor Activity Against Taxane- and Epothilone-Resistant Cancer Cells.
Trigili, C., Barasoain, I., Sanchez-Murcia, P.A., Bargsten, K., Redondo-Horcajo, M., Nogales, A., Gardner, N.M., Meyer, A., Naylor, G.J., Gomez-Rubio, E., Gago, F., Steinmetz, M.O., Paterson, I., Prota, A.E., Diaz, J.F.(2016) ACS Omega 1: 1192-1204
- PubMed: 30023505
- DOI: https://doi.org/10.1021/acsomega.6b00317
- Primary Citation of Related Structures:
5MF4 - PubMed Abstract:
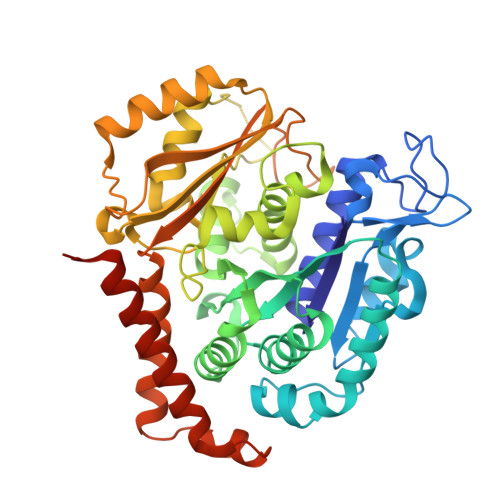
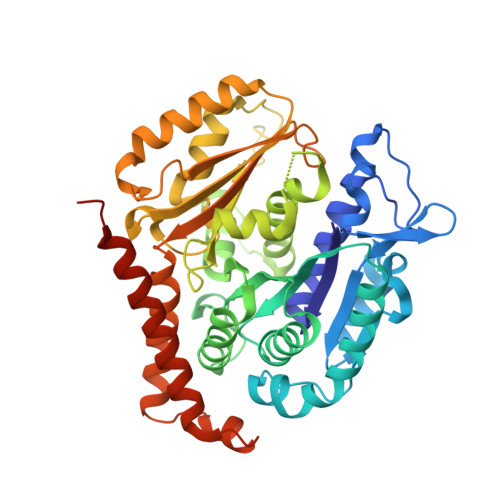
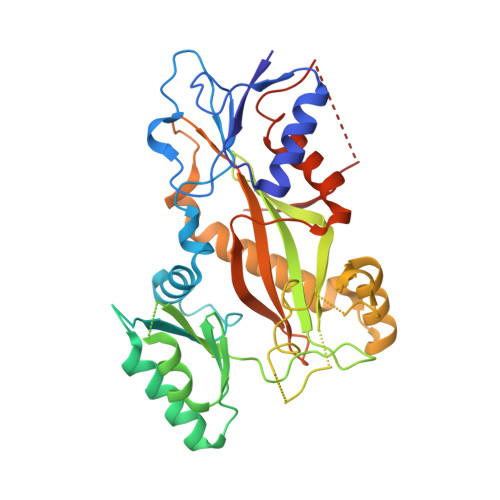
A combined biochemical, structural, and cell biology characterization of dictyostatin is described, which enables an improved understanding of the structural determinants responsible for the high-affinity binding of this anticancer agent to the taxane site in microtubules (MTs). The study reveals that this macrolide is highly optimized for MT binding and that only a few of the structural modifications featured in a library of synthetic analogues resulted in small gains in binding affinity. The high efficiency of the dictyostatin chemotype in overcoming various kinds of clinically relevant resistance mechanisms highlights its potential for therapeutic development for the treatment of drug-resistant tumors. A structural explanation is advanced to account for the synergy observed between dictyostatin and taxanes on the basis of their differential effects on the MT lattice. The X-ray crystal structure of a tubulin-dictyostatin complex and additional molecular modeling have allowed the rationalization of the structure-activity relationships for a set of synthetic dictyostatin analogues, including the highly active hybrid 12 with discodermolide. Altogether, the work reported here is anticipated to facilitate the improved design and synthesis of more efficacious dictyostatin analogues and hybrids with other MT-stabilizing agents.
- Chemical and Physical Biology, Centro de Investigaciones Biológicas, CSIC, Ramiro de Maeztu 9, E-28040 Madrid, Spain.
Organizational Affiliation: