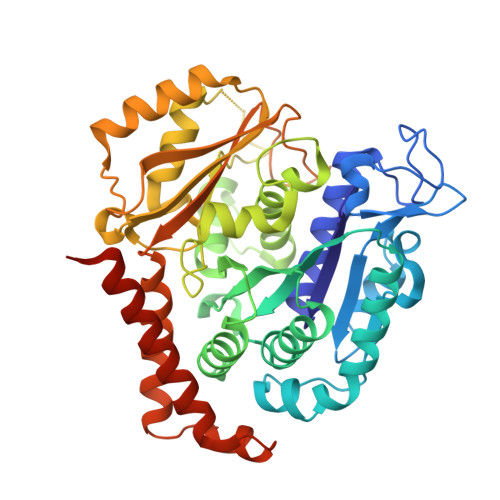
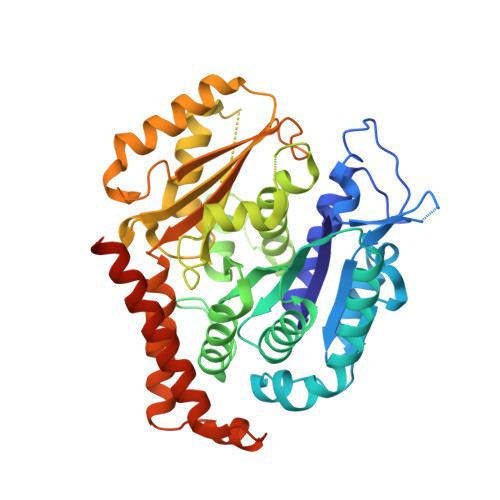
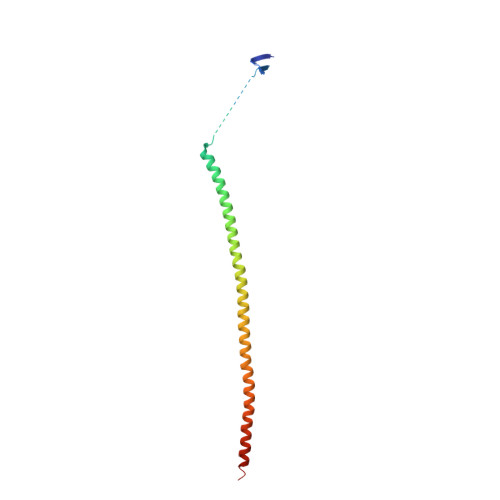
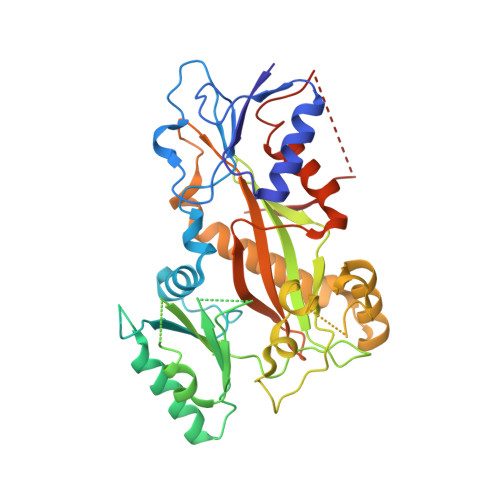
Triazolopyrimidines Are Microtubule-Stabilizing Agents that Bind the Vinca Inhibitor Site of Tubulin.
Saez-Calvo, G., Sharma, A., Balaguer, F.A., Barasoain, I., Rodriguez-Salarichs, J., Olieric, N., Munoz-Hernandez, H., Berbis, M.A., Wendeborn, S., Penalva, M.A., Matesanz, R., Canales, A., Prota, A.E., Jimenez-Barbero, J., Andreu, J.M., Lamberth, C., Steinmetz, M.O., Diaz, J.F.(2017) Cell Chem Biol 24: 737-750.e6
- PubMed: 28579361
- DOI: https://doi.org/10.1016/j.chembiol.2017.05.016
- Primary Citation of Related Structures:
5NJH - PubMed Abstract:
Microtubule-targeting agents (MTAs) are some of the clinically most successful anti-cancer drugs. Unfortunately, instances of multidrug resistances to MTA have been reported, which highlights the need for developing MTAs with different mechanistic properties. One less explored class of MTAs are [1,2,4]triazolo[1,5-a]pyrimidines (TPs). These cytotoxic compounds are microtubule-stabilizing agents that inexplicably bind to vinblastine binding site on tubulin, which is typically targeted by microtubule-destabilizing agents. Here we used cellular, biochemical, and structural biology approaches to address this apparent discrepancy. Our results establish TPs as vinca-site microtubule-stabilizing agents that promote longitudinal tubulin contacts in microtubules, in contrast to classical microtubule-stabilizing agents that primarily promote lateral contacts. Additionally we observe that TPs studied here are not affected by p-glycoprotein overexpression, and suggest that TPs are promising ligands against multidrug-resistant cancer cells.
Organizational Affiliation:
Department of Physical and Chemical Biology, Centro de Investigaciones Biológicas, Consejo Superior de Investigaciones Científicas, Ramiro de Maeztu 9, 28040 Madrid, Spain.