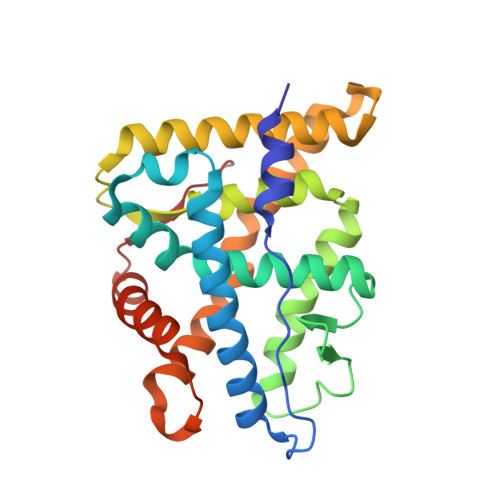
First High-Resolution Crystal Structures of the Glucocorticoid Receptor Ligand-Binding Domain-Peroxisome Proliferator-ActivatedgammaCoactivator 1-alphaComplex with Endogenous and Synthetic Glucocorticoids.
Liu, X., Wang, Y., Ortlund, E.A.(2019) Mol Pharmacol 96: 408-417
- PubMed: 31391291
- DOI: https://doi.org/10.1124/mol.119.116806
- Primary Citation of Related Structures:
6NWK, 6NWL - PubMed Abstract:
Both synthetic and endogenous glucocorticoids are important pharmaceutic drugs known to bind to the ligand-binding domain (LBD) of glucocorticoid receptor (GR), a member of the nuclear receptor (NR) superfamily. Ligand binding induces conformational changes within GR, resulting in subsequent DNA binding and differential coregulator recruitment, ultimately activating or repressing target gene expression. One of the most crucial coregulators is peroxisome proliferator-activated γ coactivator 1- α (PGC1 α ), which acts to regulate energy metabolism by directly interacting with GR to modulate gene expression. However, the mechanisms through which PGC1 α senses GR conformation to drive transcription are not completely known. Here, an ancestral variant of the GR (AncGR2) LBD was used as a tool to produce stable protein for biochemical and structural studies. PGC1 α is found to interact more tightly and form a more stable complex with AncGR2 LBD than nuclear receptor coactivator 2. We report the first high-resolution X-ray crystal structures of AncGR2 LBD in complex with PGC1 α and dexamethasone (DEX) or hydrocortisone (HCY). Structural analyses reveal how distinct steroid drugs bind to GR with different affinities by unique hydrogen bonds and hydrophobic interactions. Important charge clamps are formed between the activation function 2 and PGC1 α to mediate their specific interactions. These interactions lead to a high level of protection from hydrogen-deuterium exchange at the coregulator interaction site and strong intramolecular allosteric communication to ligand binding site. This is the first structure detailing the GR-PGC1 α interaction providing a foundation for future design of specific therapeutic agents targeting these critical metabolic regulators. SIGNIFICANCE STATEMENT: High-resolution structures of AncGR2 LBD bound to DEX and HCY in complex with PGC1α reveal the molecular mechanism of PGC1α binding to AncGR2 LBD as well as the distinct affinities between DEX and HCY binding. Identifying the structural mechanisms that drive drug affinity is of pharmacologic interest to the glucocorticoid receptor field as an avenue to guide future drug design targeting GR-PGC1α signaling, which plays a crucial role in controlling hepatic glucose output.
Organizational Affiliation:
Department of Biochemistry, Emory University School of Medicine, Atlanta Georgia (X.L., Y.W., E.A.O.) and College of Life Sciences, Qingdao University, Qingdao, People's Republic of China (Y.W.).