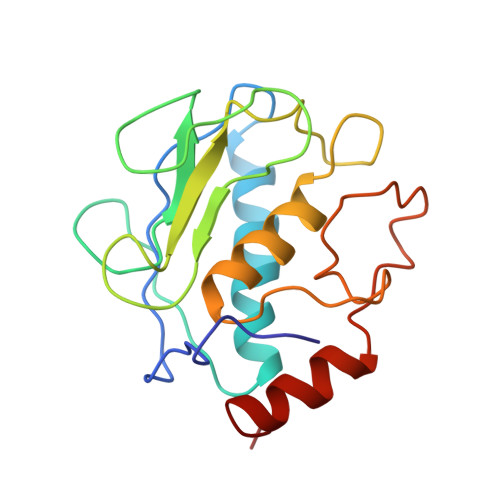
Solution structures of stromelysin complexed to thiadiazole inhibitors.
Stockman, B.J., Waldon, D.J., Gates, J.A., Scahill, T.A., Kloosterman, D.A., Mizsak, S.A., Jacobsen, E.J., Belonga, K.L., Mitchell, M.A., Mao, B., Petke, J.D., Goodman, L., Powers, E.A., Ledbetter, S.R., Kaytes, P.S., Vogeli, G., Marshall, V.P., Petzold, G.L., Poorman, R.A.(1998) Protein Sci 7: 2281-2286
- PubMed: 9827994
- DOI: https://doi.org/10.1002/pro.5560071105
- Primary Citation of Related Structures:
3USN - PubMed Abstract:
Unregulated or overexpressed matrix metalloproteinases (MMPs), including stromelysin, collagenase, and gelatinase. have been implicated in several pathological conditions including arthritis and cancer. Small-molecule MMP inhibitors may have therapeutic value in the treatment of these diseases. In this regard, the solution structures of two stromelysin/ inhibitor complexes have been investigated using 1H, 13C, and 15N NMR spectroscopy. Both-inhibitors are members of a novel class of matrix metalloproteinase inhibitor that contain a thiadiazole group and that interact with stromelysin in a manner distinct from other classes of inhibitors. The inhibitors coordinate the catalytic zinc atom through their exocyclic sulfur atom, with the remainder of the ligand extending into the S1-S3 side of the active site. The binding of inhibitor containing a protonated or fluorinated aromatic ring was investigated using 1H and 19F NMR spectroscopy. The fluorinated ring was found to have a reduced ring-flip rate compared to the protonated version. A strong, coplanar interaction between the fluorinated ring of the inhibitor and the aromatic ring of Tyr155 is proposed to account for the reduced ring-flip rate and for the increase in binding affinity observed for the fluorinated inhibitor compared to the protonated inhibitor. Binding interactions observed for the thiadiazole class of ligands have implications for the design of matrix metalloproteinase inhibitors.
Organizational Affiliation:
Structural, Analytical & Medicinal Chemistry, Pharmacia & Upjohn, Kalamazoo, Michigan 49001, USA. brian.j.stockman@am.pnu.com