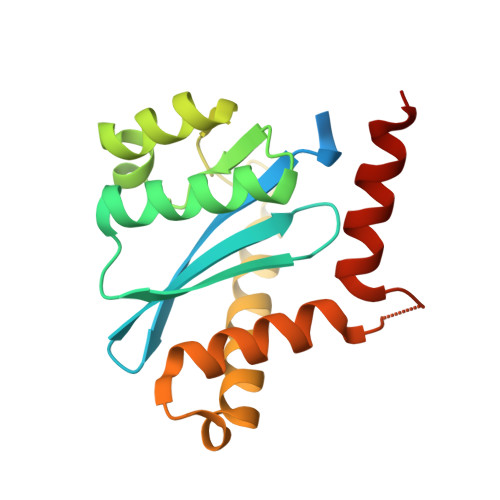
Interrogating HIV Integrase for Compounds that Bind- a Sampl Challenge.
Peat, T.S., Dolezal, O., Newman, J., Mobley, D., Deadman, J.J.(2014) J Comput Aided Mol Des 28: 347
- PubMed: 24532034
- DOI: https://doi.org/10.1007/s10822-014-9721-7
- Primary Citation of Related Structures:
4CE9, 4CEA, 4CEB, 4CEC, 4CED, 4CEE, 4CEF, 4CEO, 4CEQ, 4CER, 4CES, 4CEZ, 4CF0, 4CF1, 4CF2, 4CF8, 4CF9, 4CFA, 4CFB, 4CFC, 4CFD, 4CGD, 4CGF, 4CGG, 4CGH, 4CGI, 4CGJ, 4CHN, 4CHO, 4CHP, 4CHQ, 4CHY, 4CHZ, 4CIE, 4CIF, 4CIG, 4CJ3, 4CJ4, 4CJ5, 4CJE, 4CJF, 4CJK, 4CJL, 4CJP, 4CJQ, 4CJR, 4CJS, 4CJT, 4CJU, 4CJV - PubMed Abstract:
Tremendous gains and novel methods are often developed when people are challenged to do something new or difficult. This process is enhanced when people compete against each other-this can be seen in sport as well as in science and technology (e.g. the space race). The SAMPL challenges, like the CASP challenges, aim to challenge modellers and software developers to develop new ways of looking at molecular interactions so the community as a whole can progress in the accurate prediction of these interactions. In order for this challenge to occur, data must be supplied so the prospective test can be done. We have supplied unpublished data related to a drug discovery program run several years ago on HIV integrase for the SAMPL4 challenge. This paper describes the methods used to obtain these data and the chemistry involved.
Organizational Affiliation:
CSIRO, Parkville, VIC, Australia, tompeat@hotmail.com.