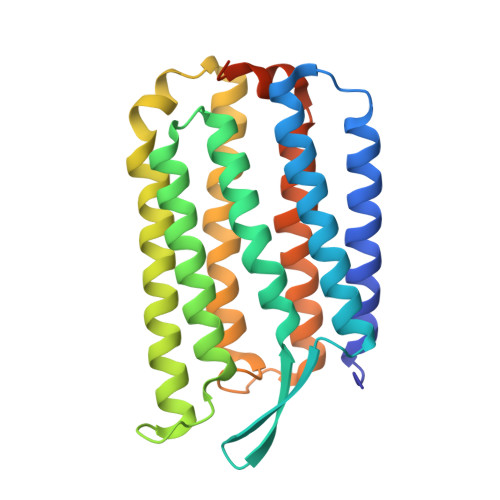
Crystal Structure of Escherichia coli-Expressed Haloarcula marismortui Bacteriorhodopsin I in the Trimeric Form.
Shevchenko, V., Gushchin, I., Polovinkin, V., Round, E., Borshchevskiy, V., Utrobin, P., Popov, A., Balandin, T., Buldt, G., Gordeliy, V.(2014) PLoS One 9: e112873-e112873
- PubMed: 25479443
- DOI: https://doi.org/10.1371/journal.pone.0112873
- Primary Citation of Related Structures:
4PXK - PubMed Abstract:
Bacteriorhodopsins are a large family of seven-helical transmembrane proteins that function as light-driven proton pumps. Here, we present the crystal structure of a new member of the family, Haloarcula marismortui bacteriorhodopsin I (HmBRI) D94N mutant, at the resolution of 2.5 Å. While the HmBRI retinal-binding pocket and proton donor site are similar to those of other archaeal proton pumps, its proton release region is extended and contains additional water molecules. The protein's fold is reinforced by three novel inter-helical hydrogen bonds, two of which result from double substitutions relative to Halobacterium salinarum bacteriorhodopsin and other similar proteins. Despite the expression in Escherichia coli and consequent absence of native lipids, the protein assembles as a trimer in crystals. The unique extended loop between the helices D and E of HmBRI makes contacts with the adjacent protomer and appears to stabilize the interface. Many lipidic hydrophobic tail groups are discernible in the membrane region, and their positions are similar to those of archaeal isoprenoid lipids in the crystals of other proton pumps, isolated from native or native-like sources. All these features might explain the HmBRI properties and establish the protein as a novel model for the microbial rhodopsin proton pumping studies.
Organizational Affiliation:
Institute of Complex Systems (ICS-6) Structural Biochemistry, Research Centre Jülich GmbH, Jülich, Germany; Laboratory for advanced studies of membrane proteins, Moscow institute of physics and technology, Dolgoprudniy, Russia.