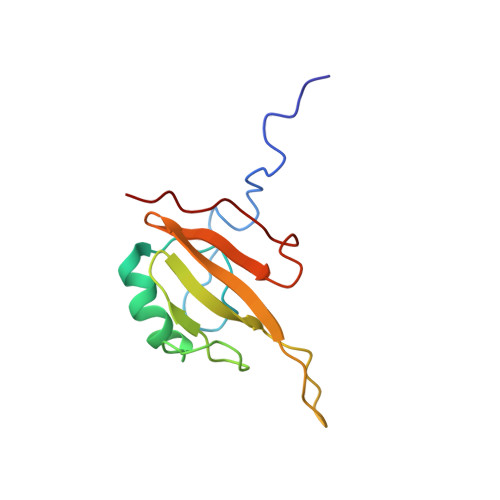
Induced conformational changes activate the peptidoglycan synthase PBP1B.
Egan, A.J.F., Maya-Martinez, R., Ayala, I., Bougault, C.M., Banzhaf, M., Breukink, E., Vollmer, W., Simorre, J.P.(2018) Mol Microbiol 110: 335-356
- PubMed: 30044025
- DOI: https://doi.org/10.1111/mmi.14082
- Primary Citation of Related Structures:
6FZK, 6G5R, 6G5S - PubMed Abstract:
Bacteria surround their cytoplasmic membrane with an essential, stress-bearing peptidoglycan (PG) layer consisting of glycan chains linked by short peptides into a mesh-like structure. Growing and dividing cells expand their PG layer using inner-membrane anchored PG synthases, including Penicillin-binding proteins (PBPs), which participate in dynamic protein complexes to facilitate cell wall growth. In Escherichia coli, and presumably other Gram-negative bacteria, growth of the mainly single layered PG is regulated by outer membrane-anchored lipoproteins. The lipoprotein LpoB is required to activate PBP1B, which is a major, bi-functional PG synthase with glycan chain polymerising (glycosyltransferase) and peptide cross-linking (transpeptidase) activities. In this work we show how the binding of LpoB to the regulatory UB2H domain of PBP1B activates both activities. Binding induces structural changes in the UB2H domain, which transduce to the two catalytic domains by distinct allosteric pathways. We also show how an additional regulator protein, CpoB, is able to selectively modulate the TPase activation by LpoB without interfering with GTase activation.
Organizational Affiliation:
The Centre for Bacterial Cell Biology, Institute for Cell and Molecular Biosciences, Newcastle University, Richardson Road, Newcastle upon Tyne, NE2 4AX, UK.