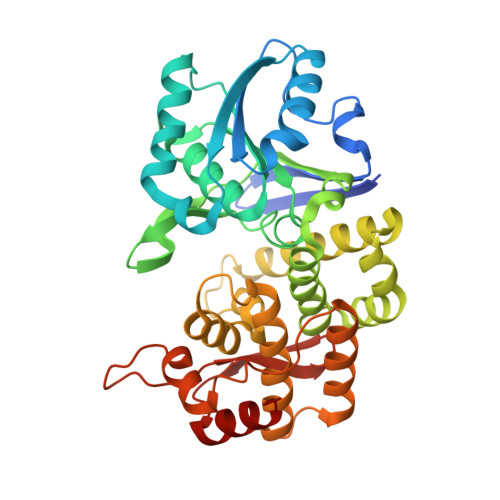
Structural and Biochemical Analyses Reveal that Chlorogenic Acid Inhibits the Shikimate Pathway.
Neetu, N., Katiki, M., Dev, A., Gaur, S., Tomar, S., Kumar, P.(2020) J Bacteriol 202
- PubMed: 32661075
- DOI: https://doi.org/10.1128/JB.00248-20
- Primary Citation of Related Structures:
6LK2, 6LLA - PubMed Abstract:
Chlorogenic acid (CGA) is a phenolic compound with well-known antibacterial properties against pathogens. In this study, structural and biochemical characterization was used to show the inhibitory role of CGA against the enzyme of the shikimate pathway, a well-characterized drug target in several pathogens. Here, we report the crystal structures of dehydroquinate synthase (DHQS), the second enzyme of the shikimate pathway, from Providencia alcalifaciens ( Pa DHQS), in binary complex with NAD and ternary complex with NAD and CGA. Structural analyses reveal that CGA occupies the substrate position in the active site of Pa DHQS, which disables domain movements, leaving the enzyme in an open and catalysis-incompetent state. The binding analyses by isothermal titration calorimetry (ITC) and surface plasmon resonance (SPR) show that CGA binds to Pa DHQS with K D (equilibrium dissociation constant) values of 6.3 μM and 0.5 μM, respectively. In vitro e nzyme inhibition studies show that CGA inhibits Pa DHQS with a K i of 235 ± 21 μM, while it inhibits the growth of Providencia alcalifaciens , Moraxella catarrhalis , Staphylococcus aureus , and Escherichia coli with MIC values of 60 to 100 μM. In the presence of aromatic amino acids supplied externally, CGA does not show the toxic effect. These results, along with the observations of the inhibition of the 3-deoxy-d-arabino-heptulosonate-7-phosphate (DAHP) regulatory domain by CGA in our previous study, suggest that CGA binds to shikimate pathway enzymes with high affinity and inhibits their catalysis and can be further exploited for designing novel drug-like molecules. IMPORTANCE The shikimate pathway is an attractive target for the development of herbicides and antimicrobial agents, as it is essential in plants, bacteria, and apicomplexan parasites but absent in humans. The enzymes of shikimate pathway are conserved among bacteria. Thus, the inhibitors of the shikimate pathway act on wide range of pathogens. We have identified that chlorogenic acid targets the enzymes of the shikimate pathway. The crystal structure of dehydroquinate synthase, the second enzyme of the pathway, in complex with chlorogenic acid and enzymatic inhibition studies explains the mechanism of inhibition of chlorogenic acid. These results suggest that chlorogenic acid has a good chemical scaffold and have important implications for its further development as a potent inhibitor of shikimate pathway enzymes.
Organizational Affiliation:
Department of Biotechnology, Indian Institute of Technology Roorkee, Roorkee, Uttarakhand, India.