Fragment based drug design - Small chemical changes of fragments effecting big changes in binding
Oebbeke, M., Wienen-Schmidt, B., Heine, A., Klebe, G.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| cAMP-dependent protein kinase catalytic subunit alpha | 353 | Cricetulus griseus | Mutation(s): 0 Gene Names: PRKACA EC: 2.7.11.11 | 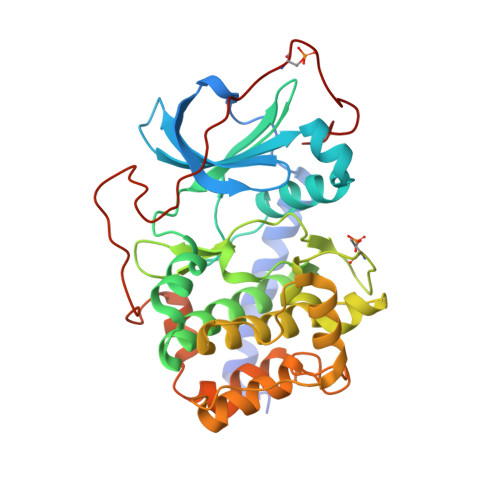 | |
UniProt | |||||
Find proteins for P25321 (Cricetulus griseus) Explore P25321 Go to UniProtKB: P25321 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P25321 | ||||
Sequence AnnotationsExpand | |||||
| |||||
Find similar proteins by: Sequence | 3D Structure
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| cAMP-dependent protein kinase inhibitor alpha | 20 | Mus musculus | Mutation(s): 0 | 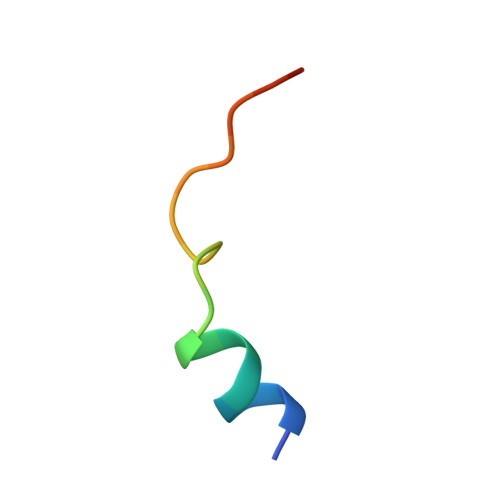 | |
UniProt & NIH Common Fund Data Resources | |||||
Find proteins for P63248 (Mus musculus) Explore P63248 Go to UniProtKB: P63248 | |||||
IMPC: MGI:104747 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P63248 | ||||
Sequence AnnotationsExpand | |||||
| |||||
| Ligands 3 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| ADE (Subject of Investigation/LOI) Query on ADE | C [auth A] | ADENINE C5 H5 N5 GFFGJBXGBJISGV-UHFFFAOYSA-N |  | ||
| MPD Query on MPD | E [auth A] | (4S)-2-METHYL-2,4-PENTANEDIOL C6 H14 O2 SVTBMSDMJJWYQN-YFKPBYRVSA-N |  | ||
| MRD Query on MRD | D [auth A], F [auth B] | (4R)-2-METHYLPENTANE-2,4-DIOL C6 H14 O2 SVTBMSDMJJWYQN-RXMQYKEDSA-N |  | ||
| Modified Residues 2 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Type | Formula | 2D Diagram | Parent |
| SEP Query on SEP | A | L-PEPTIDE LINKING | C3 H8 N O6 P |  | SER |
| TPO Query on TPO | A | L-PEPTIDE LINKING | C4 H10 N O6 P |  | THR |
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 72.926 | α = 90 |
| b = 75.122 | β = 90 |
| c = 80.792 | γ = 90 |
| Software Name | Purpose |
|---|---|
| PHENIX | refinement |
| PHASER | phasing |
| Coot | model building |
| XDS | data reduction |
| XDS | data scaling |