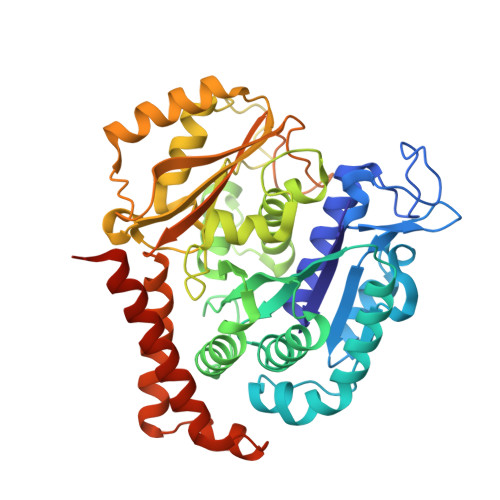
Reversible binding of the anticancer drug KXO1 (tirbanibulin) to the colchicine-binding site of beta-tubulin explains KXO1's low clinical toxicity.
Niu, L., Yang, J., Yan, W., Yu, Y., Zheng, Y., Ye, H., Chen, Q., Chen, L.(2019) J Biol Chem 294: 18099-18108
- PubMed: 31628188
- DOI: https://doi.org/10.1074/jbc.RA119.010732
- Primary Citation of Related Structures:
6KNZ - PubMed Abstract:
KXO1 (tirbanibulin or KX2-391) is as a non-ATP-competitive inhibitor of SRC proto-oncogene nonreceptor tyrosine kinase (SRC) and is being clinically investigated for the management of various cancers and actinic keratosis. Recently, KXO1 has also been shown to strongly inhibit tubulin. Interestingly, unlike conventional tubulin-targeting drugs, KXO1 has exhibited low toxicity in preclinical and clinical studies, but the reason for this remains elusive, as are the KXO1-binding site and other details of the interaction of KXO1 with tubulin. Here, cell-based experiments revealed that KXO1 induces tubulin depolymerization and G 2 /M phase cell cycle arrest at low nanomolar concentrations, similar to colchicine, used as a positive control. Results from biochemical experiments, including an N , N -ethylenebis(iodoacetamide) competition assay, disclosed that KXO1 binds to the colchicine-binding site on β-tubulin, further confirmed by the crystal structure of the tubulin-KXO1 complex at 2.5-Å resolution. A high-quality electron density map of the crystallographic data enabled us to unambiguously determine the position and orientation of KXO1 in the colchicine-binding site, revealing the detailed interactions between KXO1 and tubulin. We also found that KXO1 binds reversibly to purified tubulin, induces a totally reversible cellular effect (G 2 /M cell cycle arrest), and possesses no cellular toxicity 5 days after drug washout, explaining KXO1's low toxicity. In summary, we show that KXO1 binds to the colchicine-binding site of tubulin and resolved the crystal structure of the tubulin-KXO1 complex. Importantly, KXO1's reversible binding to tubulin explains its clinically low toxicity, an insight that could guide further clinical applications of KXO1.
Organizational Affiliation:
State Key Laboratory of Biotherapy and Cancer Center, West China Hospital of Sichuan University and Collaborative Innovation Center of Biotherapy and Cancer, Chengdu 610041, China.