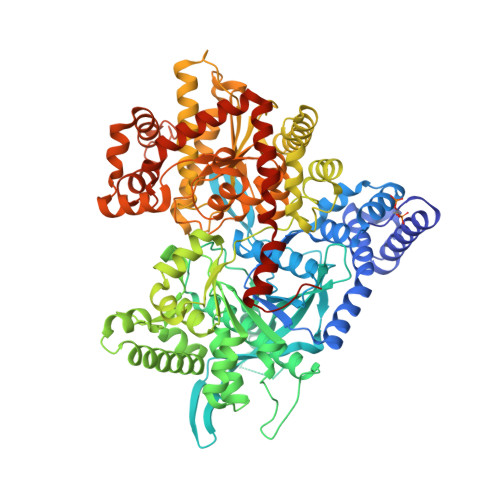
Anthranilimide based glycogen phosphorylase inhibitors for the treatment of type 2 diabetes. Part 3: X-ray crystallographic characterization, core and urea optimization and in vivo efficacy.
Thomson, S.A., Banker, P., Bickett, D.M., Boucheron, J.A., Carter, H.L., Clancy, D.C., Cooper, J.P., Dickerson, S.H., Garrido, D.M., Nolte, R.T., Peat, A.J., Sheckler, L.R., Sparks, S.M., Tavares, F.X., Wang, L., Wang, T.Y., Weiel, J.E.(2009) Bioorg Med Chem Lett 19: 1177-1182
- PubMed: 19138846
- DOI: https://doi.org/10.1016/j.bmcl.2008.12.085
- Primary Citation of Related Structures:
3DD1, 3DDS, 3DDW - PubMed Abstract:
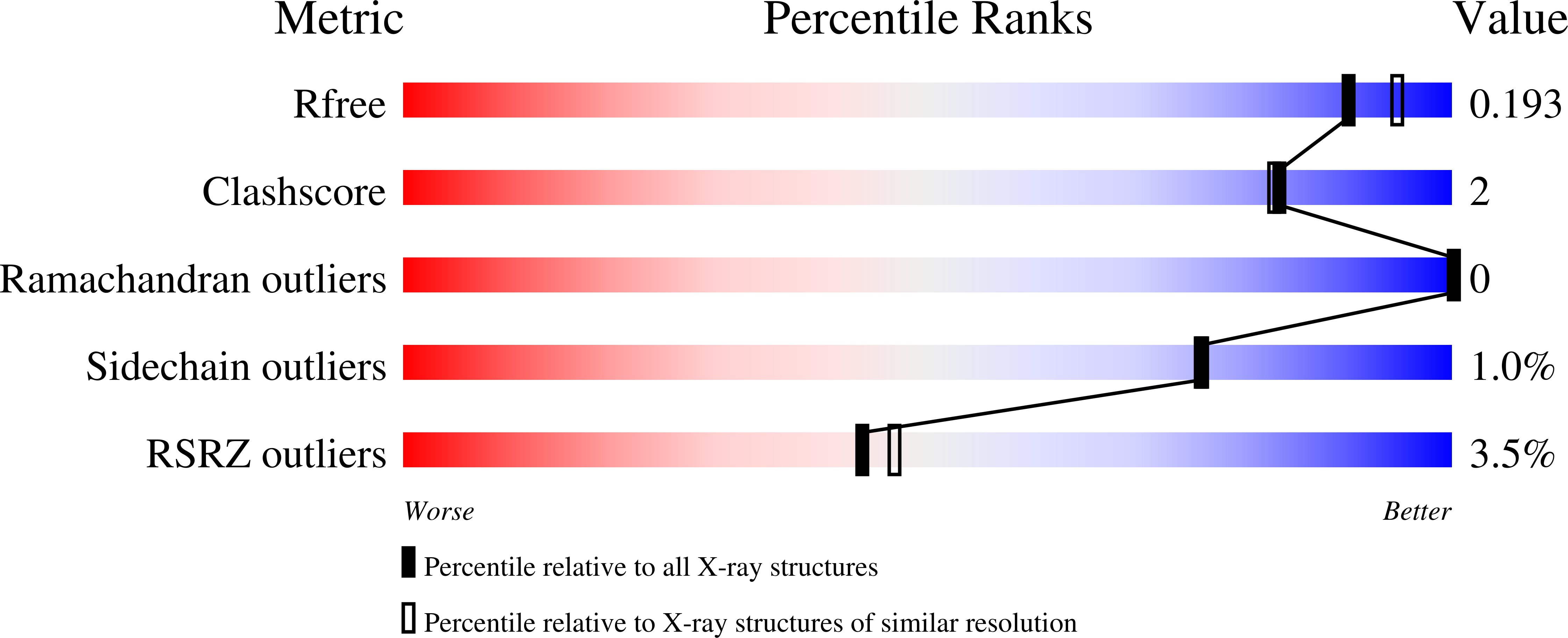
Key binding interactions of the anthranilimide based glycogen phosphorylase a (GPa) inhibitor 2 from X-ray crystallography studies are described. This series of compounds bind to the AMP site of GP. Using the binding information the core and the phenyl urea moieties were optimized. This work culminated in the identification of compounds with single nanomolar potency as well as in vivo efficacy in a diabetic model.
Organizational Affiliation:
Metabolic Center for Excellence in Drug Discovery, GlaxoSmithKline, 5 Moore Drive, PO Box 13398, Research Triangle Park, NC 27705, USA. stephen.a.thomson@gsk.com